Genome #10: Udgam of India - Unraveling a genetic khichdi
Where did the modern Indians come from?
I moved to the US for a PhD in the Fall of 2014. Adapting to new environments wasn’t new to me. I had moved places in India and had spent some time abroad for an internship. I, however, distinctly remember one question that I would get asked during small talks or in the social hour after our department seminars - “Where are you from”? The answer would be simple - “I am from India”. “Where in India?” - “Bombay, you probably would have heard about it”.
The part about me being from Bombay is only partially true. I was born in the small town of Jhakra in Samastipur, Bihar, grew up in the smallish town of Rawatbhata near Kota in Rajasthan and spent my teenage years in Bombay. I am sure this trajectory of growing up in different places is not unique to me, but the simplification of saying “I am from Bombay” was driven in part by my love for the city and in part by its recognizability. “Bombay”, that answer, came with some sense of completeness - everyone knows about it. The most usual response would be, “My best friend is from Bombay,” which I would follow up with, “It is the best city on earth!”. Every time I repeated one or the other version of my "I am from Bombay" answer confidentially, I also felt equally puzzled inside - where did I really come from? Who were my ancestors? Were they always in India? Were they permanently settled in Bihar?
Where did I come from?
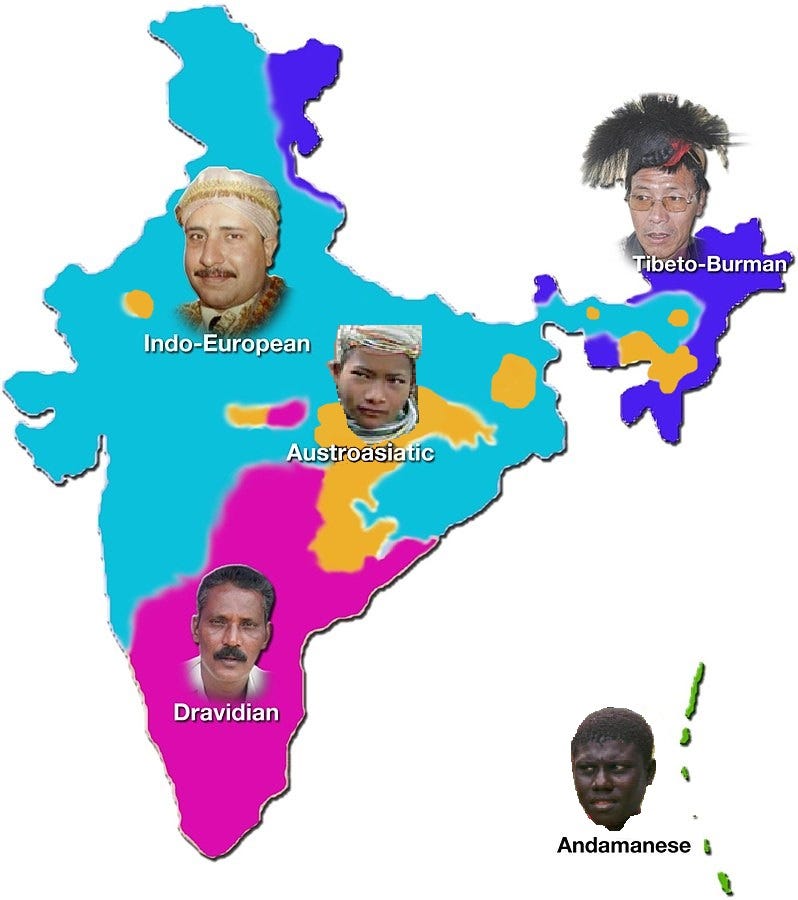
Where did I come from? Were my ancestors all ‘Indian’? Why does this question matter at all? In 2015, when I started thinking about the question sincerely, the question was purely curiosity-driven. I genuinely wanted to know where my udgam (source) was. There are multiple ways to ask this question. First, I could return to my family records to trace my ancestors' origins. Unfortunately, these records are nonexistent; no one in my family knows anything beyond the two generations preceding them. An alternate way to narrow my search would be to focus on linguistics. I speak Hindi, while my parents speak Maithili, both members of the Indo-European language family (Figure 1), which has been primarily in use in north India. I could narrow my search by looking into how these languages evolved - did Maithili originate in Bihar initially, and did all my ancestors speak the same language? I am not a linguist, so following my roots linguistically would have been challenging. The third and easiest way was to rely on genetics - my training was in Chemical Engineering, but I had come to the US for a Ph.D. in Computational Biology, where I was learning about a new concept in Biology every day (ten years later, I still do) that I also see being applied in real life - from sequencing our DNA to gene therapies. I will explain how computational genetics help us trace our roots in a bit. But this third way was easiest because companies like 23andMe and Ancestry.com have commercially available kits that profile your DNA, looking for specific changes at specific places that provide a good indication of where you and your ancestors came from. So, I ordered a swab kit from 23andMe and gave away my genetic history of myself and people directly related to me for the ‘greater good.’
My 23andMe results did not reveal anything striking - my ancestral DNA belongs to “North India” (Figure 2). Though unsurprising, the answer brought a bit more completeness to my usual answer - “I come from Bombay, but I am ancestrally from Bihar”. Besides this, the information I obtained from 23andMe did not add anything specific to my knowledge (I remember using a third-party service to get finer-grained results), and by the end of 2015, I had put the question to rest.
How does our DNA explain who we are?
Genetics can be intimidating if you haven’t studied it earlier. But the basics necessary to understand the rest of the story are straightforward.
All tissues in the body are ultimately made of cells. Each cell has a nucleus that stores most of your “genetic information” - the DNA. We inherit our “genetic code” or the DNA from our parents packed in 23 pairs of chromosomes inside the cell's nucleus. All cells have exactly the same DNA - so the DNA in your hair is the same as the DNA in your toes. DNA is composed of four bases or nucleotides: Adenine (A), Guanine (G), Thymine (T), and Cytosine (C). Chromosomes vary in length, and their total length is 3 billion bases. So, our genetic code is a unique sequence of 3 billion bases made of A/C/T/Gs (Figure 3).

The idea behind using genetics to figure out our roots is very ingenious yet very simple. It relies on the fact that as humans reproduce, the mother and father each pass half of their DNA to the child (this also holds for most vertebrate life forms on the earth). So, on average, each half of the 23 pairs of chromosomes is contributed by each of the parents except the Y chromosome. All females have two copies of the X chromosome, but males have one copy of the X and one copy of the Y chromosome. The remaining chromosomes are named chromosomes 1-22. The Y chromosome is passed in entirety from the father to the son. Between your genome and mine, each 3 billion letters long, there are 3 million letters that are different on average, which makes you you and me me. The higher these differences are on any DNA segment, the longer it has been since the two shared a common ancestor (Figure 4). By looking at these differences (and similarities), we can figure out who is our “genetic neighbor” and “genetic ancestor”. This is also the concept used by companies like Ancestry.com and 23andMe.com to identify people belonging to the same family tree. This is enabled by decades of progress in sequencing technology, which allows us to determine at base-level resolution the identity of each of the 3 billion base pairs. However, you do not need to determine all 3 billion base pairs to answer ancestry-or-related questions. Sometimes, a few thousand of bases can provide accurate answers.

The Indian subcontinent genetics - a thali?
Shashi Tharoor, in his book “The Elephant, The Tiger and The Cell Phone: Reflections on India” calls India a thali: If America is a melting pot, then to me India is a thali, a selection of sumptuous dishes in different bowls. Each tastes different and does not necessarily mix with the next, but they belong together on the same plate, and they complement each other in making the meal a satisfying repast.
While Shashi Tharoor’s statement had nothing to do with genetics, it is worth considering whether the diversity we notice today in terms of what people look like, their languages, and their distinct traditions mean they are also different genetically. To understand how similar or dissimilar the current data Indians are, David Reich, an American geneticist whose group has done seminal work in the field of genetics of ancient humans and their migration, collaborated with Dr. Lalji Singh and Dr. Kumarswamy Thangaraj at the Centre for Cellular and Molecular Biology (CCMB) in Hyderabad in 2009. They profiled genetic mutations in hundreds of current-day individuals belonging to 20+ distinct groups (spanning multiple states, spoken languages, and caste/religion).
Just studying sequence from the pool of current Indians would only tell us how similar (or dissimilar) they are. But to ask a broader question on where the current Indians likely came from, we have to look also outside India. The research study analyzed genetic data of current-day Indians along with non-Indians, comprising European Americans, Georgians, Iranians, basque, and Han-Chinese, using a mathematical technique of principal component analysis (PCA). PCA finds a combination of single nucleotide changes in the DNA that are most informative in describing the overall genetic distance between individuals in an unsupervised fashion (which means that it does not care about which individuals came from which group or subgroup). PCA representation on a two-dimensional graph showed that the Indian samples spread out along a gradient line (“a cline”), with the Indo-European speakers on one extreme and the Dravidian speakers on the other extreme (Figure 5B). The west Eurasian people lay close to the Indo-European speakers while the Dravidian speakers, which lie farthest away from west Eurasians, are close to the Andamanese and Austro-asiatic speakers, yet the latter two groups are genetically very distinct.

What does the “cline” on the PCA plot tell us? The data points from the Dravidian-speaking subgroup (“south” Indians) lie very far from the West Eurasians on the cline, and the Indo-European language speakers (“north” Indians) lie all along the cline. A simple explanation would be that the current-day Indians are a genetic mixture of two different populations - like the simplest khichdi made of two components besides water: rice and dal (pulses). So genetically, we are sumptuous khichdis currently residing in different bowls of the Indian thali.
So if we are a result of a mixture of two components - what are these original ingredients? It is possible that the Europeans and current-day Indians belong to the same arm of the family tree that branched out from the ancestors of the Chinese population, such as the Han-Chinese subgroup. To test this, we can look at differences in DNA sequences between the European and current-day Indians and then check what these differences look like in the Chinese population. The Chinese subgroup had more similarities to the Indian DNA sequences than they do with the Europeans, implying that Europeans and Indians likely did not descend from a homogeneous ancestral population. The next obvious possibility is that the Indians and Chinese could have descended from the same common ancestral population. But this possibility also does not hold groups because when the Europeans are compared to all the Indians, they have more similarities than the Chinese.
The DNA sequences observed in current-day Indians, on average, are intermediate between those in Europeans and East Asians. The only way these intermediate similarities can arise is if the ancestral populations of Europeans and Central Asians mixed with distinctly related ancestors of the East Asians. The only exceptions are the people from the island of Andamans - who are isolated descendants of an East-Asian-related population.
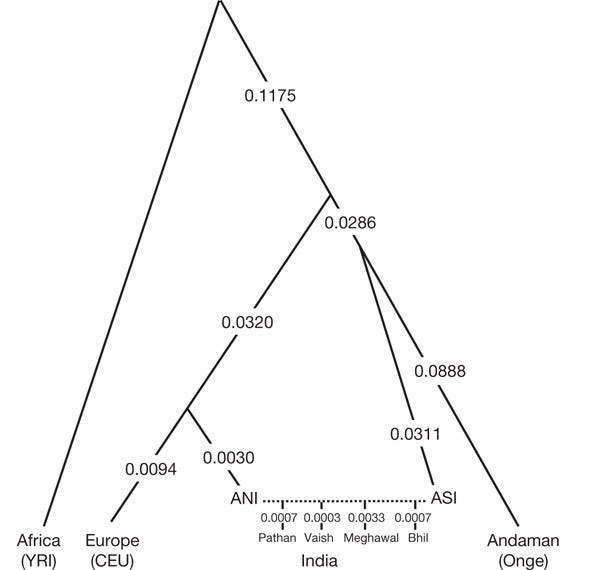
Based on evidence in two papers published in 2009 and 2013, the authors concluded (padding it softly to avoid any political issues arising from the pre-conceived and incorrect notion of racial purity of Indians), that all modern-day Indians are descendants of two highly differentiated populations: Ancestral North Indians (ANI) and Ancestral South Indians (ASI) (Figure 6). ANI population is related to Europeans and central Asians, while the ASI descended from a population unrelated to any present-day population outside India. ANI and ASI mixed (interbred) in India extensively; hence, everyone in India is a genetic mixture that started happening almost 5,000 years before the present (Figure 7). Any claims of genetic purity are outright false, even for ASI and ANI populations.

The past of caste - a latent threat for rare genetic disorders
“Jo kabhi nahin jaati, usiko jati kehte hain” .
The thing that never goes, that thing is called caste. – From the movie ‘Swades’
How have the genetic marks of history not been eroded even after thousands of years? One of the most distinctive features of our society, though not exclusive, is ‘caste’ - the system of social stratification that determines who will marry whom. Though our constitution prohibits discrimination based on caste, caste continues to shape who Indians marry today.
A sociological definition of caste (jati) is “A stratified system of categorization based on a status conferred at birth (ascribed status) based on descent, in which individuals do not have mobility due to custom or law.” The stratification manifests in India primarily through endogamy - preventing people from marrying outside their castes. And hence “Jo kabhi nahin jaati, usiko jati kehte hain”.
When David Reich’s group analyzed the genetic data of current-day Indians spanning multiple jatis, they made a remarkable observation. Compared to European groups separated by the same geographical distance, the genetic makeup of two jatis in India is at least three times distinct! This effect cannot be explained by the difference in ANI ancestry percentage (over ASI) or the difference in the geographical location where these samples were profiled. What, then, explains the increased degree of genetic homogeneity between members of the same jati?
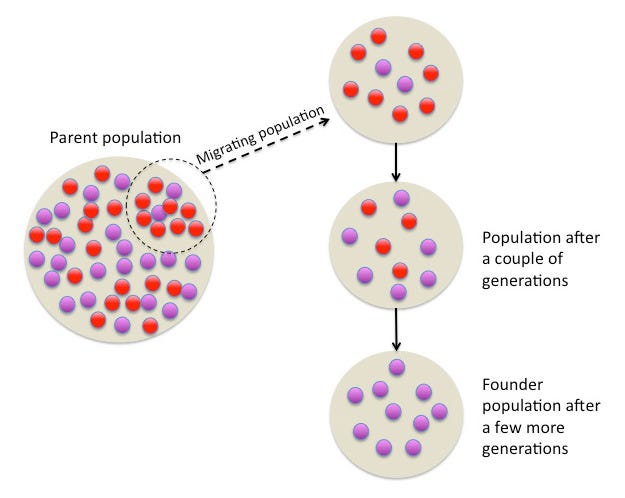
The Indian jatis today may be a product of “population bottlenecks.” Imagine a few individuals having many offspring and then having many more offspring yet remaining genetically isolated from people surrounding them due to social barriers - induced by endogamy. For example, it is estimated that around 4,000 years ago, the Finnish population went through a strong “population bottleneck,” resulting in exceptionally low diversity in the Y chromosome - likely reflective of the survival of just two ancestral male lineages. Similarly, a large fraction of the ancestry of today’s Ashkenazi Jews is attributable to population bottlenecks or founder effects (Figure 8) between AD 1100 and AD 1400.
The hallmark signs of population bottlenecks are visible in the Indian genetic data spanning multiple jatis - these manifest as long stretches of almost identical DNA between two individuals from the same jati. The only possibility in this case of observing near identical long stretches of DNA is if the individuals descended from the same ancestors in the last few years (who also carried this segment of DNA). Around one-third of Indian jatis face population bottlenecks or founder effects as strong as the Ashkenazi Jews or the Finns (Figure 9).

What can we learn from our genetic khichdi?
There are vast implications of founder effects. For example, in certain parts of southern India, before undergoing surgery, you are asked your caste. However bizarre that may sound, there is a scientific reason behind it - they are often looking for if you belong to the Vysya community - a regional group associated with traders and business people primarily living around Andhra Pradesh and Telangana. Anecdotally, the anesthesiologists know that Vysya community people can have a fatal reaction to common muscle relaxants, so they need a different combination of drugs to prevent any reaction.
So why do people from the Vysya community react differently to muscle relaxants? Founder effects - Vysya ancestors lived not in geographic isolation but in densely populated Andhra Pradesh. Despite geographical proximity, Vysya community like most jatis in India follows strict social isolation by enforcing endogamy rules, which have been followed by subsequent generations. People of the Vysya community are known to have genetically lower levels of butyrylcholinesterase, a protein that helps break down components of standard muscle relaxants. Lower levels of this protein in Vysya community arise because of a mutation originally found in an ancestor of the Vysya community. How has the mutation been passed on so long over generations? Because both the parents have exactly the same mutation at the same site - prolonged endogamy forces the child to inherit these mutations (and keep passing them on to their offspring).
In this example of the suffering of one community, there is a wonderful opportunity -given the high rate of endogamy and rare genetic disorders arising from ‘homozygous mutations’ (where both father and mother have a certain mutation inherited from an ancestor). To catalog such rare genetic disorders, we need to sequence the genomes of individuals from the same jati and identify jatis with strong population effects. In one such study, David Reich’s group characterized that the founder event strength is at least three times higher in the Vysya community, but some communities like Kapu and Banyas have much stronger founder effects. The authors did not characterize the rare diseases the other groups had, but a catalog of such diseases was prepared with the help of local doctors and midwives who anecdotally have much more information about the diseases and malformations in babies to understand how these diseases can be managed.
The impact of understanding the impact of rare mutations would impact hundreds of thousands of people. A catalog of such diseases and the genes causing them would enable us to understand how changes in sequence can increase or decrease our chances of carrying certain diseases. For example, Tay-Sachs disease is a rare genetic disorder caused by “homozygous mutations” that affect babies and young children. Children with Tay-Sachs have lower levels of an enzyme that breaks down fatty substances, which ultimately affects the function of neurons in the brain. Tay-Sachs disease used to be common in the Ashkenazi Jewish community, but several genetic testing organizations have helped control its occurrence. How do you control a rare genetic disorder? Ashkenazi Jewish community also has a system of arranged marriages similar to India. Genetic counseling organizations test teenagers to see if they are “carriers” of the mutation known to cause Tay-Sachs and other rare genetic diseases. If they are carriers, the matchmakers do not introduce them to other “carriers”. A small first step in India would be replicating this model at scale. This would first require thoroughly characterizing rare genetic diseases in the Indian context and the associated mutations.
Every time I think of my udgam, of the city streets in Bombay or the small town in Bihar, I am reminded not just of my immediate past but of tens of thousands of years of history. A history that is as much mine as it is of every individual who calls this vast subcontinent home. We can learn a lot from our genetic history, providing us with a more informed future for ourselves and future generations.
References
Tamang R, Thangaraj K. Genomic view on the peopling of India. Investigative genetics. 2012 Dec;3:1-9.
Reich D, Thangaraj K, Patterson N, Price AL, Singh L. Reconstructing Indian population history. Nature. 2009 Sep 24;461(7263):489-94.
Moorjani P, Thangaraj K, Patterson N, Lipson M, Loh PR, Govindaraj P, Berger B, Reich D, Singh L. Genetic evidence for recent population mixture in India. The American Journal of Human Genetics. 2013 Sep 5;93(3):422-38.
Reich D. Who we are and how we got here: Ancient DNA and the new science of the human past. Oxford University Press; 2018 Mar 23.
Narasimhan VM, Patterson N, Moorjani P, Rohland N, Bernardos R, Mallick S, Lazaridis I, Nakatsuka N, Olalde I, Lipson M, Kim AM. The formation of human populations in South and Central Asia. Science. 2019 Sep 6;365(6457):eaat7487.
Nakatsuka N, Moorjani P, Rai N, Sarkar B, Tandon A, Patterson N, Bhavani GS, Girisha KM, Mustak MS, Srinivasan S, Kaushik A. The promise of discovering population-specific disease-associated genes in South Asia. Nature genetics. 2017 Sep 1;49(9):1403-7.







Appreciate the depth of research in your posts Saket - pls keep writing!